qsm_sb: Fast quantitative susceptibility mapping¶
Contents
- 1. Print qsm_sb information
- 2. Setting model parameters
- 2.a. Create qsm_sb object
- 2.b. Set protocol and options
- 2.b.1 Set protocol the CLI way
- 2.b.2 Set protocol and options the GUI way
- 3. Fit MRI data
- 3.a. Load input data
- 3.b. Execute fitting process
- 3.c. Display FitResults
- 3.d. Save fit results
- 3.e. Re-use or share fit configuration files
- 4. Simulations
- 4.a. Single Voxel Curve
- 4.b. Sensitivity Analysis
- 5. Notes
- 5.a. Notes specific to qsm_sb
- 5.a.1 BIDS
- 5.a.2 Note-1
- 5.a.3 Note-2
- 5.a.4 Note-3
- 5.b. Generic notes
- 5.b.1. Batch friendly option and protocol conventions
- 5.b.2 Parallelization:
- 6. Citations
% This m-file has been automatically generated using qMRgenBatch(qsm_sb) % for publishing documentation. % Command Line Interface (CLI) is well-suited for automatization % purposes and Octave. % % Please execute this m-file section by section to get familiar with batch % processing for qsm_sb on CLI. % % Demo files are downloaded into qsm_sb_data folder. % % Written by: Agah Karakuzu, 2017 % ==============================================================================
1. Print qsm_sb information
qMRinfo('qsm_sb');
qsm_sb: Fast quantitative susceptibility mapping Assumptions: Type/number of outputs will depend on the selected options. (1) Case - Split-Bregman: i) W/ magnitude weighting: chiSBM, chiL2M, chiL2, unwrappedPhase, maskOut ii) W/O magnitude weighting: chiSM, chiL2, unwrappedPhase, maskOut (2) Case - L2 Regularization: i) W/ magnitude weighting: chiL2M, chiL2, unwrappedPhase, maskOut ii) W/O magnitude weighting: chiL2, unwrappedPhase, maskOut (3) Case - No Regularization: i) Magnitude weighting is not enabled: nfm, unwrappedPhase, maskOut Inputs: PhaseGRE 3D GRE acquisition. |Wrapped phase image| (MagnGRE) 3D GRE acquisition. |Magnitude part of the image| (OPTIONAL) Mask Brain extraction mask. Outputs: chiSBM Susceptibility map created using Split-Bregman method with magnitude weighting chiSB Susceptibility map created using Split-Bregman method without magnitude weighting. chiL2M Susceptibility map created using L2 regularization with magnitude weighting chiL2 Susceptibility map created using L2 regularization without magnitude weighting nfm Susceptibility map created without regularization unwrappedPhase Unwrapped phase image using Laplacian-based method maskOut Binary mask (maskSharp, gradientMask or same as the input) Options: Derivative direction Direction of the derivation - forward - backward SHARP Filtering Sophisticated harmonic artifact reduction for phase data - State: true/false - Mode: once/iterative - Padding Size: [1X3 array] - Magnitude Weighting: on/off L1-Regularization Apply L1-regularization - State: true/false - Reoptimize parameters: true/false - Lambda-L1: [double] - L1-Range: [1X2 array] L2-Regularization Apply L2-regularization - State: true/false - Reoptimize parameters: true/false - Lambda-L2: [double] - L2-Range: [1X2 array] Split-Bregman Apply Split-Bregman method - State: true/false - Reoptimize parameters: Authors: Agah Karakuzu, 2018 References: Please cite the following if you use this module: Bilgic et al. (2014), Fast quantitative susceptibility mapping with L1-regularization and automatic parameter selection. Magn. Reson. Med., 72: 1444-1459. doi:10.1002/mrm.25029 In addition to citing the package: Karakuzu A., Boudreau M., Duval T.,Boshkovski T., Leppert I.R., Cabana J.F., Gagnon I., Beliveau P., Pike G.B., Cohen-Adad J., Stikov N. (2020), qMRLab: Quantitative MRI analysis, under one umbrella doi: 10.21105/joss.02343 Reference page in Doc Center doc qsm_sb
2. Setting model parameters
2.a. Create qsm_sb object
Model = qsm_sb;
2.b. Set protocol and options
Protocol: MRI acquisition parameters that are accounted for by the respective model.
For example: TE, TR, FA FieldStrength. The assigned protocol values are subjected to a sanity check to ensure that they are in agreement with the data attributes.
Options: Fitting preferences that are left at user's discretion.
For example: linear fit, exponential fit, drop first echo.
2.b.1 Set protocol the CLI way
If you are using Octave, or would like to serialize your operations any without GUI involvement, you can assign protocol directly in CLI:
xDim = 0.6; yDim = 0.6; zDim = 0.6; Model.Prot.Resolution.Mat = [ xDim yDim zDim ];
TE = 0.0081; Model.Prot.Timing.Mat = [ TE ];
FieldStrength = 3; CentralFreq = 42.58; Model.Prot.Magnetization.Mat = [ FieldStrength CentralFreq ];
See the generic notes section below for further information.
2.b.2 Set protocol and options the GUI way
The following command opens a panel to set protocol and options (if GUI is available to the user):
Model = Custom_OptionsGUI(Model);

If available, you need to close this panel for the remaining of the script to proceed.
Using this panel, you can save qMRLab protocol files that can be used in both interfaces. See the generic notes section below for details.
3. Fit MRI data
3.a. Load input data
This section shows how you can load data into a(n) qsm_sb object.
- At the CLI level, qMRLab accepts structs containing (double) data in the fields named in accordance with a qMRLab model.
See the generic notes section below for BIDS compatible wrappers and scalable
qMRLab workflows.
% |- qsm_sb object needs 3 data input(s) to be assigned: % |- PhaseGRE % |- MagnGRE % |- Mask data = struct(); % PhaseGRE.mat contains [40 40 40] data. load('qsm_sb_data/PhaseGRE.mat'); % MagnGRE.mat contains [40 40 40] data. load('qsm_sb_data/MagnGRE.mat'); % Mask.mat contains [40 40 40] data. load('qsm_sb_data/Mask.mat'); data.PhaseGRE= double(PhaseGRE); data.MagnGRE= double(MagnGRE); data.Mask= double(Mask);
3.b. Execute fitting process
This section will fit the loaded data.
FitResults = FitData(data,Model,0);
Visit the generic notes section below for instructions to accelerate fitting by
parallelization using ParFitData.
3.c. Display FitResults
You can display the current outputs by:
qMRshowOutput(FitResults,data,Model);
A representative fit curve will be plotted if available.
To render images in this page, we will load the fit results that had been saved before. You can skip the following code block;
% Load FitResults that comes with the example dataset. FitResults_old = load('FitResults/FitResults.mat'); qMRshowOutput(FitResults_old,data,Model);
3.d. Save fit results
Outputs can be saved as *.nii.(gz) if NIfTI inputs are available:
% Generic function call to save nifti outputs FitResultsSave_nii(FitResults, 'reference/nifti/file.nii.(gz)');
If not, FitResults.mat file can be saved. This file contains all the outputs as workspace variables:
% Generic function call to save FitResults.mat
FitResultsSave_mat(FitResults);
FitResults.mat files can be loaded to qMRLab GUI for visualization and ROI
analyses.
The section below will be dynamically generated in accordance with the example data format (mat or nii). You can substitute FitResults_old with FitResults if you executed the fitting using example dataset for this model in section 3.b..
FitResultsSave_mat(FitResults_old);
3.e. Re-use or share fit configuration files
qMRLab's fit configuration files (qsm_sb_Demo.qmrlab.mat) store all the options and protocol in relation to the used model and the release version.
*.qmrlab.mat files can be easily shared with collaborators to allow them fit their own
data or run simulations using identical option and protocol configurations.
Model.saveObj('my_qsm_sb_config.qmrlab.mat');
4. Simulations
4.a. Single Voxel Curve
Simulates single voxel curves
Not available for the current model.
4.b. Sensitivity Analysis
Simulates sensitivity to fitted parameters
Not available for the current model.
5. Notes
5.a. Notes specific to qsm_sb
5.a.1 BIDS

|== sub-01/ |~~~~~~ anat/ |---------- sub-01_T1w.json |---------- sub-01_part-mag_T1w.nii.gz |---------- sub-01_part-phase_T1w.nii.gz | |== derivatives/ |~~~~~~ qMRLab/ |---------- dataset_description.json |~~~~~~~~~~ sub-01/anat/ |-------------- sub-01_Chimap.nii.gz |-------------- sub-01_Chimap.json |~~~~~~ pipeline/ |---------- dataset_description.json |~~~~~~~~~~ sub-01/anat/ |-------------- sub-01_label-brain_mask.nii.gz |-------------- sub-01_label-brain_mask.json
For further information, please visit BIDS qMRI Appendix.
5.a.2 Note-1
To make documentation generation and our CI tests faster for this model, we used a subportion of the data 40x40x40 in our testing environment.
Therefore , the dimensions displayed in the section 3.a of this automatically generated documentation is different than those provided to the user.
qMRgenBatch(qsm_sb)
When the command above is called from MATLAB/Octave by the user, the complete high resolution (0.6mm iso resolution) dataset (384x336x224 dimensions) will be downloaded.
5.a.3 Note-2
- L1 regularization is performed as a sub-process to Split-Bregman regularization. Therefore, L1 (lambda) reoptimization panel is enabled only when Split-Bregman is activated. The Chimap (inn ppm) calculated upon this selection are chiSB and chiL2.
- Unlike L1, L2 regularization can be run independently. The Chimap (in ppm) calculated upon this selection is chiL2.
- When the magnitude weighting optio`n is enbled, associated the output maps attains an M suffix: chiL2M or chiSBM.
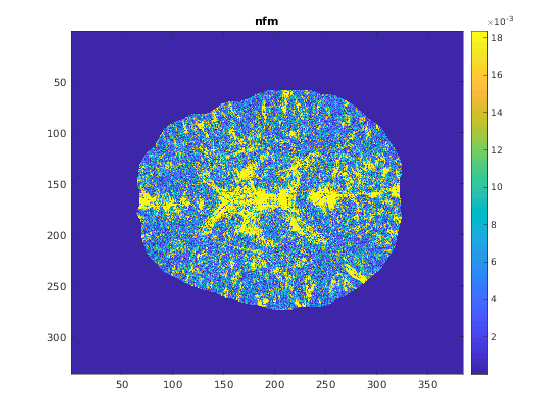
- A susceptibility map can be also generated without regularization. For this selection, the output Chimap is nfm, which is displayed as an example in this page.
- Unwrapped phase images are provided in the output: unwrappedPhase.
5.a.4 Note-3
Full dataset and the original code are available here.
5.b. Generic notes
5.b.1. Batch friendly option and protocol conventions
If you would like to load a desired set of options / protocols programatically, you can use *.qmrlab.mat files. To save a configuration from the protocol panel of qsm_sb, first open the respective panel by running the following command in your MATLAB command window (MATLAB only):
Custom_OptionsGUI(qsm_sb);
In this panel, you can arrange available options and protocols according to your needs, then click the save button to save my_qsm_sb.qmrlab.mat file. This file can be later loaded into a qsm_sb object in batch by:
Model = qsm_sb;
Model = Model.loadObj('my_qsm_sb.qmrlab.mat');
Model.loadObj('my_qsm_sb.qmrlab.mat') call won't update the fields in the Model object, unless the output is assigned to the object as shown above. This compromise on convenience is to retain Octave CLI compatibility.
If you don't have MATLAB, hence cannot access the GUI, two alternatives are available to populate options:
- Use qmrlab/mcrgui:latest Docker image to access GUI. The instructions are available here.
- Set options and protocols in CLI:
- List available option fields using tab completion in Octave's command prompt (or window)
Model = qsm_sb;
Model.option. % click the tab button on your keyboard and list the available fields.
- Assign the desired field. For example, for a mono_t2 object:
Model = mono_t2; Model.options.DropFirstEcho = true; Model.options.OffsetTerm = false;
Some option fields may be mutually exclusive or interdependent. Such cases are handled by the GUI options panel; however, not exposed to the CLI. Therefore, manual CLI options assignments may be challenging for some involved methods such as qmt_spgr or qsm_sb. If above options are not working for you and you cannot infer how to set options solely in batch, please feel free to open an issue in qMRLab and request the protocol file you need.
Similarly, in CLI, you can inspect and assign the protocols:
Model = qsm_sb;
Model.Prot. % click the tab button on your keyboard and list the available fields.
Each protocol field has two subfields of Format and Mat. The first one is a cell indicating the name of the protocol parameter (such as EchoTime (ms)) and the latter one contains the respective values (such as 30 x 1 double array containing EchoTimes).
The default Mat protocol values are set according to the example datasets served via OSF.
5.b.2 Parallelization:
The current model does not perform voxelwise fitting. Therefore, parallelization is not enabled.
6. Citations
qMRLab JOSS article
Karakuzu A., Boudreau M., Duval T.,Boshkovski T., Leppert I.R., Cabana J.F., Gagnon I., Beliveau P., Pike G.B., Cohen-Adad J., Stikov N. (2020), qMRLab: Quantitative MRI analysis, under one umbrella 10.21105/joss.02343
Reference article for qsm_sb
Quantitative MRI, under one umbrella.
NeuroPoly Lab, Montreal, Canada