amico: Accelerated Microstructure Imaging via Convex Optimization¶
Contents
- 1. Print amico information
- 2. Setting model parameters
- 2.a. Create amico object
- 2.b. Set protocol and options
- 2.b.1 Set protocol the CLI way
- 2.b.2 Set protocol and options the GUI way
- 3. Fit MRI data
- 3.a. Load input data
- 3.b. Execute fitting process
- 3.c. Display FitResults
- 3.d. Save fit results
- 3.e. Re-use or share fit configuration files
- 4. Simulations
- 4.a. Single Voxel Curve
- 4.b. Sensitivity Analysis
- 5. Notes
- 5.a. Notes specific to amico
- 5.b. Generic notes
- 5.b.1. Batch friendly option and protocol conventions
- 5.b.2 Parallelization
- 6. Citations
% This m-file has been automatically generated using qMRgenBatch(amico) % for publishing documentation. % Command Line Interface (CLI) is well-suited for automatization % purposes and Octave. % % Please execute this m-file section by section to get familiar with batch % processing for amico on CLI. % % Demo files are downloaded into amico_data folder. % % Written by: Agah Karakuzu, 2017 % ==============================================================================
1. Print amico information
qMRinfo('amico');
amico: Accelerated Microstructure Imaging via Convex Optimization Sub-module of noddi Pulse Sequence Diagram ASSUMPTIONS: Neuronal fibers model: geometry sticks (Dperp = 0) Orientation dispersion YES (Watson distribution). Note that NODDI is more robust to crossing fibers that DTI (Campbell, NIMG 2017) Permeability NO Diffusion properties: intra-axonal totally restricted diffusion coefficient (Dr) fixed by default. extra-axonal Tortuosity model. Parallel diffusivity is equal to intra-diffusivity.Perpendicular diffusivity is proportional to fiber density diffusion coefficient (Dh) Constant Inputs: DiffusionData 4D diffusion weighted dataset (Mask) Binary mask to accelerate the fitting (OPTIONAL) Outputs: di Diffusion coefficient in the restricted compartment. ficvf Fraction of water in the restricted compartment. fiso Fraction of water in the isotropic compartment (e.g. CSF/Veins) fr Fraction of restricted water in the entire voxel (e.g. intra-cellular volume fraction) fr = ficvf*(1-fiso) irfrac Fraction of isotropically restricted compartment (Dot for ex vivo model) diso (fixed) diffusion coefficient of the isotropic compartment (CSF) kappa Orientation dispersion index b0 Signal at b=0 theta angle of the fibers phi angle of the fibers Protocol: Multi-shell diffusion-weighted acquisition at least 2 non-zeros bvalues at least 5 b=0 (used to compute noise standard deviation DiffusionData Array [NbVol x 7] Gx Diffusion Gradient x Gy Diffusion Gradient y Gz Diffusion Gradient z Gnorm (T/m) Diffusion gradient magnitude Delta (s) Diffusion separation delta (s) Diffusion duration TE (s) Echo time Options: Model Model part of NODDI. Available models are: -WatsonSHStickTortIsoVIsoDot_B0 is a four model compartment used for ex-vivo datasets Example of command line usage For more examples: qMRusage(noddi) Author: Tanguy Duval References: Please cite the following if you use this module: Alessandro Daducci, Erick Canales-Rodriguez, Hui Zhang, Tim Dyrby, Daniel Alexander, Jean-Philippe Thiran, 2015. Accelerated Microstructure Imaging via Convex Optimization (AMICO) from diffusion MRI data. NeuroImage 105, pp. 32-44 Zhang, H., Schneider, T., Wheeler-Kingshott, C.A., Alexander, D.C., 2012. NODDI: practical in vivo neurite orientation dispersion and density imaging of the human brain. Neuroimage 61, 1000?1016. In addition to citing the package: Karakuzu A., Boudreau M., Duval T.,Boshkovski T., Leppert I.R., Cabana J.F., Gagnon I., Beliveau P., Pike G.B., Cohen-Adad J., Stikov N. (2020), qMRLab: Quantitative MRI analysis, under one umbrella doi: 10.21105/joss.02343 Documentation for amico doc amico
2. Setting model parameters
2.a. Create amico object
Model = amico;
2.b. Set protocol and options
Protocol: MRI acquisition parameters that are accounted for by the respective model.
For example: TE, TR, FA FieldStrength. The assigned protocol values are subjected to a sanity check to ensure that they are in agreement with the data attributes.
Options: Fitting preferences that are left at user's discretion.
For example: linear fit, exponential fit, drop first echo.
2.b.1 Set protocol the CLI way
If you are using Octave, or would like to serialize your operations any without GUI involvement, you can assign protocol directly in CLI:
Not available for the current model.
See the generic notes section below for further information.
2.b.2 Set protocol and options the GUI way
The following command opens a panel to set protocol and options (if GUI is available to the user):
Model = Custom_OptionsGUI(Model);
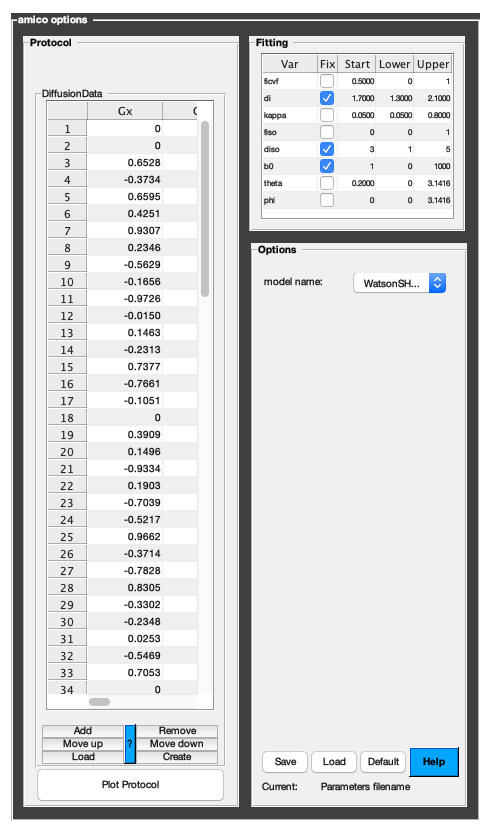
You need to close this window for the remaining of the script to proceed.
Using this panel, you can save qMRLab protocol files that can be used in both interfaces. See the generic notes section below for details.
3. Fit MRI data
3.a. Load input data
This section shows how you can load data into a(n) amico object.
- At the CLI level, qMRLab accepts structs containing (double) data in the fields named in accordance with a qMRLab model.
See the generic notes section below for BIDS compatible wrappers and scalable
qMRLab workflows.
% |- amico object needs 2 data input(s) to be assigned: % |- DiffusionData % |- Mask data = struct(); % DiffusionData.nii.gz contains [74 87 50 109] data. data.DiffusionData=double(load_nii_data('amico_data/DiffusionData.nii.gz')); % Mask.nii.gz contains [74 87 50] data. data.Mask=double(load_nii_data('amico_data/Mask.nii.gz'));
3.b. Execute fitting process
This section will fit the loaded data.
FitResults = FitData(data,Model,0);
Visit the generic notes section below for instructions to accelerate fitting by
parallelization using ParFitData.
3.c. Display FitResults
You can display the current outputs by:
qMRshowOutput(FitResults,data,Model);
A representative fit curve will be plotted if available.
To render images in this page, we will load the fit results that had been saved before. You can skip the following code block;
% Load FitResults that comes with the example dataset. FitResults_old = load('FitResults/FitResults.mat'); qMRshowOutput(FitResults_old,data,Model);
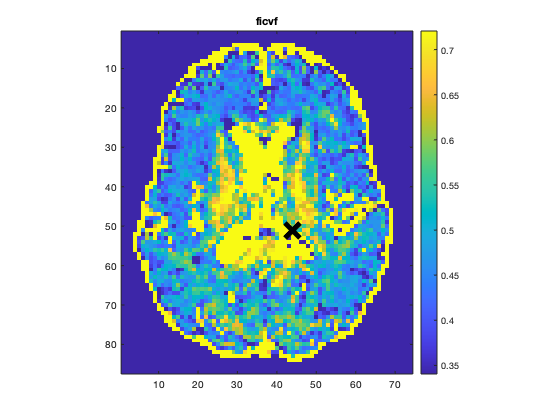
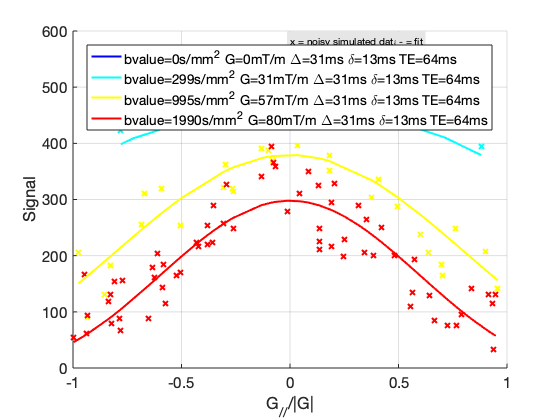
3.d. Save fit results
Outputs can be saved as *.nii.(gz) if NIfTI inputs are available:
% Generic function call to save nifti outputs FitResultsSave_nii(FitResults, 'reference/nifti/file.nii.(gz)');
If not, FitResults.mat file can be saved. This file contains all the outputs as workspace variables:
% Generic function call to save FitResults.mat
FitResultsSave_mat(FitResults);
FitResults.mat files can be loaded to qMRLab GUI for visualization and ROI
analyses.
The section below will be dynamically generated in accordance with the example data format (mat or nii). You can substitute FitResults_old with FitResults if you executed the fitting using example dataset for this model in section 3.b..
FitResultsSave_nii(FitResults_old, 'amico_data/DiffusionData.nii.gz');
Warning: Directory already exists.
3.e. Re-use or share fit configuration files
qMRLab's fit configuration files (amico_Demo.qmrlab.mat) store all the options and protocol in relation to the used model and the release version.
*.qmrlab.mat files can be easily shared with collaborators to allow them fit their own
data or run simulations using identical option and protocol configurations.
Model.saveObj('my_amico_config.qmrlab.mat');
4. Simulations
4.a. Single Voxel Curve
Simulates single voxel curves:
- Analytically generate synthetic MRI data
- Add rician noise
- Fit and plot the respective curve
x = struct; x.ficvf = 0.5; x.di = 1.7; x.kappa = 0.05; x.fiso = 0; x.diso = 3; x.b0 = 1; x.theta = 0.2; x.phi = 0; Opt.SNR = 50; % run simulation figure('Name','Single Voxel Curve Simulation'); FitResult = Model.Sim_Single_Voxel_Curve(x,Opt);
-> Precomputing rotation matrices for l_max=12: [ already computed ] -> Generating kernels with model "NODDI" for protocol "example": [ Kernels already computed. Set "doRegenerate=true" to force regeneration ] -> Resampling rotated kernels: [ ]
Error using load
Unable to read file '/Users/agah/Desktop/neuropoly/qMRLab/External/AMICO/AMICO_matlab/exports/example/kernels/NODDI/A_008.mat'. No such file or directory.
Error in AMICO_NODDI/ResampleKernels (line 118)
load( fullfile( ATOMS_path, sprintf('A_%03d.mat',progress.i) ), 'lm' );
Error in AMICO_ResampleKernels (line 48)
CONFIG.model.ResampleKernels( fullfile(AMICO_data_path,CONFIG.protocol,'kernels',CONFIG.model.id), idx_OUT, Ylm_OUT );
Error in amico/Precompute (line 142)
AMICO_ResampleKernels();
Error in amico/fit (line 156)
obj.Precompute();
Error in noddi/Sim_Single_Voxel_Curve (line 261)
FitResults = fit(obj,data);
Error in amico_batch (line 181)
FitResult = Model.Sim_Single_Voxel_Curve(x,Opt);
4.b. Sensitivity Analysis
Simulates sensitivity to fitted parameters:
- Iterate fitting parameters from lower (lb) to upper (ub) bound
- Run Sim_Single_Voxel_Curve for Nofruns times
- Compute the mean and std across runs
ficvf di kappa fiso diso b0 theta phi
OptTable.st = [0.5 1.7 0.05 0 3 1 0.2 0]; % nominal values OptTable.fx = [0 1 1 1 1 1 1 1]; %vary ficvf... OptTable.lb = [0 1.3 0.05 0 1 0 0 0]; %...from 0 OptTable.ub = [1 2.1 0.8 1 5 1e+03 3.1 3.1]; %...to 1 Opt.SNR = 50; Opt.Nofrun = 5; % run simulation SimResults = Model.Sim_Sensitivity_Analysis(OptTable,Opt); figure('Name','Sensitivity Analysis'); SimVaryPlot(SimResults, 'ficvf' ,'ficvf' );
5. Notes
5.a. Notes specific to amico
Not provided.
5.b. Generic notes
5.b.1. Batch friendly option and protocol conventions
If you would like to load a desired set of options/|protocols| programatically, you can use *.qmrlab.mat files. To save a configuration from the protocol panel of amico, first open the respective panel by running the following command in your MATLAB command window (MATLAB only):
Custom_OptionsGUI(amico);
In this panel, you can arrange available options and protocols according to your needs, then click the save button to save my_amico.qmrlab.mat file. This file can be later loaded into a amico object in batch by:
Model = amico;
Model = Model.loadObj('my_amico.qmrlab.mat');
Model.loadObj('my_amico.qmrlab.mat') call won't update the fields in the Model object, unless the output is assigned to the object as shown above. This compromise on convenience is to retain Octave CLI compatibility.
If you don't have MATLAB, hence cannot access the GUI, two alternatives are available to populate options:
- Use qmrlab/mcrgui:latest Docker image to access GUI. The instructions are available here.
- Set options and protocols in CLI:
- List available option fields using tab completion in Octave's command prompt (or window)
Model = amico;
Model.option. % click the tab button on your keyboard and list the available fields.
- Assign the desired field. For example, for a mono_t2 object:
Model = mono_t2; Model.options.DropFirstEcho = true; Model.options.OffsetTerm = false;
Some option fields may be mutually exclusive or interdependent. Such cases are handled by the GUI options panel; however, not exposed to the CLI. Therefore, manual CLI options assignments may be challenging for some involved methods such as qmt_spgr or qsm_sb. If above options are not working for you and you cannot infer how to set options solely in batch, please feel free to open an issue in qMRLab and request the protocol file you need.
Similarly, in CLI, you can inspect and assign the protocols:
Model = amico;
Model.Prot. % click the tab button on your keyboard and list the available fields.
Each protocol field has two subfields of Format and Mat. The first one is a cell indicating the name of the protocol parameter (such as EchoTime (ms)) and the latter one contains the respective values (such as 30 x 1 double array containing EchoTimes).
The default Mat protocol values are set according to the example datasets served via OSF.
5.b.2 Parallelization
Beginning from release 2.5.0, you can accelerate fitting for the voxelwise models using parallelization.
Available in MATLAB only. Requires parallel processing toolbox.
In CLI, you can perform parallel fitting by:
parpool(); FitResults = ParFitData(data,Model);
If a parpool exists, the ParFitData will use it. If not, a new pool will be created using the local profile. By default, ParFitData saves outputs automatically every 5 minutes. You can disable this feature by:
FitResults = ParFitData(data, Model, 'AutosaveEnabled', false);
Alternatively, you can change the autosave interval (min 1 min) by:
FitResults = ParFitData(data,Model,'AutoSaveInterval',10);
If something went wrong during the fitting (e.g. your computer had to be restarted), you can recover the autosaved data by:
FitResults = ParFitData(data,Model,'RecoverDirectory','/ParFitTempResults_*/folder/from/the/previous/session');
GUI users will be prompted a question about whether they would like to use parallelization after clicking the Fit Data button, if the conditions are met. When called from GUI, ParFitData will be run with default options:
- Save temporary results every 5 minutes or whenever a chunk has finished processing
- Split data into chunks with a granularity factor of 3
- Do not remove temporary fit results upon completion
For further information:
help ParFitData
The default parallelization options can be changed in the preferences.json file located at the root qMRLab directory.
6. Citations
qMRLab JOSS article
Karakuzu A., Boudreau M., Duval T.,Boshkovski T., Leppert I.R., Cabana J.F., Gagnon I., Beliveau P., Pike G.B., Cohen-Adad J., Stikov N. (2020), qMRLab: Quantitative MRI analysis, under one umbrella 10.21105/joss.02343
Reference article for amico
Quantitative MRI, under one umbrella.
NeuroPoly Lab, Montreal, Canada